Liu Lab Research
The Liu lab bridges multiple disciplines, and operates at the intersection of clinical oncology, cancer biology, and computational biology, developing computational approaches to dissect therapeutic resistance and susceptibility to chemo-, targeted-, and immuno-therapies in patient tumors. A focus is on development of rigorous methodologies using statistics and machine-learning techniques applied to specific clinical contexts to discover biologically relevant molecular correlates of response/resistance (for novel therapeutic targets and rational combination therapies) and build clinically relevant predictive models of therapeutic response/resistance. Examples of past work and collaborations include
(1) development of predictive models of response to PD-1 immune checkpoint blockade in metastatic melanoma integrating clinical, genomic, and transcriptomic features (Liu, Schilling et al Nat Medicine 2019)
(2) validation of mutations in ERCC2 (gene necessary for nucleotide excision DNA repair) as marker of susceptibility to cisplatin-based chemotherapy in muscle-invasive bladder cancer, leading to clinical trials in progress (Liu et al JAMA Onc 2016; Li et al Clin Canc Res 2019)
(3) discovering a novel class of chromatin-modulating gene mutations as drivers of oncogenesis in a standardized meta-analysis of over 1000 prostate cancers (Armenia*, Wankowic*, Liu* et al Nat Gen 2018) *co-first author
(4) de novo discovery of a cisplatin-induced mutational signature in pre- and post-cisplatin treated bladder cancers, later validated in platinum-treated lung and ovarian cancers (Liu et al Nat Comm 2017)
(5) in the same cohort, discovery of intratumoral heterogeneity and multiple gene mutations as molecular correlates of survival and therapeutic resistance (Liu et al Nat Comm 2017)
(6) dissection of immunological correlates to a signature of mismatch repair deficiency in prostate cancer (Nava Rodrigues*, Rescigno*, Liu* et al JCI 2018) *co-first author
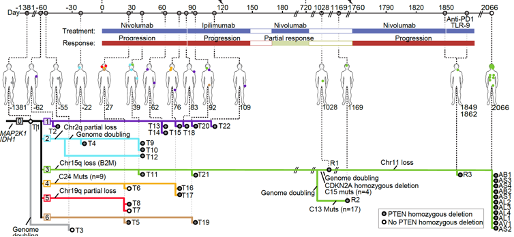
(7) elucidating evolution of therapeutic resistance and tumor heterogeneity from 37 tumor samples from a melanoma delayed exceptional responder to sequential immune checkpoint blockade collected over 6 years (Liu et al, under review)
(8) discovery of subtype-specific enrichment for secondary driver genes in genomic classes of melanoma and a DNA-repair deficiency mutational signature in a subset of melanomas in a standardized meta-analysis of over 1000 melanomas (Conway et al Nat Genet 2020)
(9) discovery of inactivating mutations in FBXW7 as a novel mechanism of PD-1 immune checkpoint blockade resistance (Gstalder, Liu et al Cancer Discov 2020)
Two broad areas of inquiry exist in the lab:
1. Biomarker discovery and development of parsimonious, biologically-informed predictive models utilizing statistical and machine learning approaches and integrating clinical and molecular data to guide clinical management.
2. Mapping of clinical tumor heterogeneity and evolution of resistance and metastasis, using longitudinally collected tumor samples from patients over time, treatments, development of resistance, and from different metastatic sites to develop insight into drivers of resistance and metastasis.
(1) development of predictive models of response to PD-1 immune checkpoint blockade in metastatic melanoma integrating clinical, genomic, and transcriptomic features (Liu, Schilling et al Nat Medicine 2019)
(2) validation of mutations in ERCC2 (gene necessary for nucleotide excision DNA repair) as marker of susceptibility to cisplatin-based chemotherapy in muscle-invasive bladder cancer, leading to clinical trials in progress (Liu et al JAMA Onc 2016; Li et al Clin Canc Res 2019)
(3) discovering a novel class of chromatin-modulating gene mutations as drivers of oncogenesis in a standardized meta-analysis of over 1000 prostate cancers (Armenia*, Wankowic*, Liu* et al Nat Gen 2018) *co-first author
(4) de novo discovery of a cisplatin-induced mutational signature in pre- and post-cisplatin treated bladder cancers, later validated in platinum-treated lung and ovarian cancers (Liu et al Nat Comm 2017)
(5) in the same cohort, discovery of intratumoral heterogeneity and multiple gene mutations as molecular correlates of survival and therapeutic resistance (Liu et al Nat Comm 2017)
(6) dissection of immunological correlates to a signature of mismatch repair deficiency in prostate cancer (Nava Rodrigues*, Rescigno*, Liu* et al JCI 2018) *co-first author
(7) elucidating evolution of therapeutic resistance and tumor heterogeneity from 37 tumor samples from a melanoma delayed exceptional responder to sequential immune checkpoint blockade collected over 6 years (Liu et al, under review)
(8) discovery of subtype-specific enrichment for secondary driver genes in genomic classes of melanoma and a DNA-repair deficiency mutational signature in a subset of melanomas in a standardized meta-analysis of over 1000 melanomas (Conway et al Nat Genet 2020)
(9) discovery of inactivating mutations in FBXW7 as a novel mechanism of PD-1 immune checkpoint blockade resistance (Gstalder, Liu et al Cancer Discov 2020)
Two broad areas of inquiry exist in the lab:
1. Biomarker discovery and development of parsimonious, biologically-informed predictive models utilizing statistical and machine learning approaches and integrating clinical and molecular data to guide clinical management.
2. Mapping of clinical tumor heterogeneity and evolution of resistance and metastasis, using longitudinally collected tumor samples from patients over time, treatments, development of resistance, and from different metastatic sites to develop insight into drivers of resistance and metastasis.
HOME | RESEARCH | PEOPLE | PUBLICATIONS | SUPPORT | CONTACT
©2019 Liu Lab at Dana-Farber Cancer Institute
©2019 Liu Lab at Dana-Farber Cancer Institute